Abstract
The asymmetric segregation of cell-fate determinants and the generation of daughter cells of different sizes rely on the correct orientation and position of the mitotic spindle. In the Drosophila embryo, the determinant Prospero is localized basally and is segregated equally to daughters of similar cell size during epidermal cell division. In contrast, during neuroblast division Prospero is segregated asymmetrically to the smaller daughter cell. This simple switch between symmetric and asymmetric segregation is achieved by changing the orientation of cell division: neural cells divide in a plane perpendicular to that of epidermoblast division. Here, by labelling mitotic spindles in living Drosophila embryos, we show that neuroblast spindles are initially formed in the same axis as epidermal cells, but rotate before cell division. We find that daughter cells of different sizes arise because the spindle itself becomes asymmetric at anaphase: apical microtubules elongate, basal microtubules shorten, and the midbody moves basally until it is positioned asymmetrically between the two spindle poles. This observation contradicts the widely held hypothesis that the cleavage furrow is always placed midway between the two centrosomes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Horvitz, H. R. & Herskowitz, I. Mechanisms in asymmetric cell division: two Bs or not two Bs, that is the question. Cell 68, 237–255 (1992).
Rappaport, R. Cytokinesis in Animal Cells (Cambridge Univ. Press, Cambridge, 1996).
Strome, S. Determination of cleavage planes. Cell 72, 3–6 (1993).
Rappaport, R. Establishment of the mechanism of cytokinesis in animal cells. Int. Rev. Cytol. 105, 245–281 (1986).
Albertson, D. Formation of the first cleavage spindle in nematode embryos. Dev. Biol. 101, 61–72 ( 1984).
Conklin, E. G. Effects of centrifugal force on the structure and development of the egg of Crepidula . J. Exp. Zool. 22, 311–419 (1917).
Matsuzaki, F., Ohshiro, T., Ikeshima-Kataoka, H. & Izumi, H. Miranda localises Staufen and Prospero asymmetrically in mitotic neuroblasts and epithelial cells in early Drosophila embryogenesis. Development 125, 4089–4098 (1998).
Doe, C. Q., Chu-LaGraff, Q., Wright, D. M. & Scott, M. P. The prospero gene specifies cell fates in the Drosophila central nervous system. Cell 65, 451– 465 (1991).
Vaessin, H. et al. prospero is expressed in neuronal precursors and encodes a nuclear protein that is involved in the control of axonal outgrowth in Drosophila. Cell 67, 941– 953 (1991).
Matsuzaki, F., Koisumi, K., Hama, C., Yoshioka, T. & Nabeshima, T. Cloning of the Drosophila prospero gene and its expression in ganglion mother cells. Biochem. Biophys. Res. Commun. 182, 1326–1332 ( 1992).
Spana, E. P. & Doe, C. Q. The Prospero transcription factor is asymmetrically localized to the cell cortex during neuroblast mitosis in Drosophila. Development 121, 3187– 3195 (1995).
Hyman, A. A. & White, J. G. Determination of cell division axes in the early embryogenesis of Caenorhabditis elegans. J. Cell Biol. 105, 2123–2135 ( 1987).
Rappaport, R. Role of mitotic apparatus in furrow initiation. Ann. NY Acad. Sci. 582, 15–21 ( 1990).
Brand, A. H. GFP in Drosophila. Trends Genet. 11, 324–325 (1995).
Brand, A. GFP as a cell and development marker in the Drosophila nervous system. Methods Cell Biol. 58, 165–181 (1999).
Callaini, G. & Anselmi, F. Centrosome splitting during nuclear elongation in the Drosophila embryo. Exp. Cell Res. 178, 415–425 (1988).
Karr, T. L. & Alberts, B. M. Organization of the cytoskeleton in early Drosophila embryos. J. Cell Sci. 102, 1494–1509 (1986).
Knoblich, J. A., Jan, L. Y. & Jan, Y. N. Deletion analysis of the Drosophila Inscuteable protein reveals domains of cortical localization and asymmetric localization. Curr. Biol. 9, 155–158 (1999).
Tio, M., Zavortink, M., Yang, X. & Chia, W. A functional analysis of inscuteable and its role during Dorsophila asymmetric cell divisions. J. Cell Sci. 112, 1541– 1551 (1999).
Kraut, R., Chia, W., Jan, L. Y., Jan, Y. N. & Knoblich, J. A. Role of inscuteable in orienting asymmetric cell divisions in Drosophila. Nature 383, 50–55 (1996).
Kraut, R. & Campos-Ortega, J. A. inscuteable, a neural precursor gene of Drosophila, encodes a candidate for a cytoskeletal adapter protein. Dev. Biol. 174, 65– 81 (1996).
Fire, A. et al. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806–811 (1998).
Moritz, M., Braunfeld, M. B., Sedat, J. W., Alberts, B. & Agard, D. A. Microtubule nucleation by γ-tubulin-containing rings in the centrosome. Nature 378, 638 –640 (1995).
Zheng, Y., Wong, M. L., Alberts, B. & Mitchison, T. Nucleation of microtubule assembly by a γ-tubulin-containing ring complex. Nature 378, 578–583 ( 1995).
Kellogg, D. R., Oegema, K., Raff, J., Schneider, K. & Alberts, B. M. CP60: a microtubule-associated protein that is localized to the centrosome in a cell-specific manner. Mol. Biol. Cell 6, 1673–1684 (1995).
Oegema, K., Whitfield, W. G. & Alberts, B. The cell cycle-dependent localization of the CP190 centrosomal protein is determined by the coordinate action of two separable domains. J. Cell Biol. 131, 1261–1273 (1995).
Whitfield, W. G., Chaplin, M. A., Oegema, K., Parry, H. & Glover, D. M. The 190 kDa centrosome-associated protein of Drosophila melanogaster contains four zinc finger motifs and binds to specific sites on polytene chromosomes. J. Cell Sci. 108, 3377–3387 (1995).
Kuchinke, U., Grawe, F. & Knust, E. Control of spindle orientation in Drosophila by the Par-3-related PDZ-domain protein Bazooka. Curr. Biol. 8, 1357–1365 (1998).
Doe, C. Q. Spindle orientation and asymmetric localization in Drosophila: both Inscuteable? Cell 86, 695– 697 (1996).
Shaw, S. L., Yeh, E., Maddox, P., Salmon, E. D. & Bloom, K. Astral microtubule dynamics in yeast: a microtubule-based searching mechanism for spindle orientation and nuclear migration into the bud. J. Cell Biol. 139, 985– 994 (1997).
Vallen, E. A., Scherson, T. Y., Roberts, T., van Zee, K. & Rose, M. D. Asymmetric mitotic segregation of the yeast spindle pole body. Cell 69, 505 –515 (1992).
Keating, H. H. & White, J. G. Centrosome dynamics in early embryos of Caenorhabditis elegans. J. Cell Sci. 111, 3027–3033 (1998).
Oegema, K. & Mitchison, T. J. Rappaport rules: cleavage furrow induction in animal cells. Proc. Natl Acad. Sci. USA 94, 4817–4820 (1997).
Bonaccorsi, S., Giansanti, M. G. & Gatti, M. Spindle self-organization and cytokinesis during male meiosis in asterless mutants of Drosophila melanogaster. J. Cell Sci. 142, 751–761 (1998).
Waddle, J. A., Cooper, J. A. & Waterston, R. H. Transient localized accumulation of actin in Caenorhabditis elegans blastomeres with oriented asymmetric division. Development 120, 2317–2328 ( 1994).
Skop, A. R. & White, J. G. The dynactin complex is required for cleavage plane specification in early Caenorhabditis elegans embryos. Curr. Biol. 8, 1110–1116 (1998).
Hyman, A. A. Centrosome movement in the early divisions of Caenorhabditis elegans: a cortical site determining centrosome position. J. Cell Biol. 109, 1185–1193 (1989).
Vale, R. D. & Toyoshima, Y. Y. Rotation and translocation of microtubules in vitro induced by dyneins from Tetrahymena cilia. Cell 52, 459–469 ( 1988).
Schuldt, A. et al. Miranda mediates asymmetric protein and RNA localisation in the developing nervous system. Genes Dev. 12, 1847–1857 (1998).
Brand, A. H. & Perrimon, N. Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development 118, 401–415 ( 1993).
Robertson, H. M. et al. A stable source of P-element transposase in Drosophila melanogaster. Genetics 118, 461– 470 (1988).
Therrien, M. et al. KSR, a novel protein kinase required for RAS signal transduction . Cell 83, 879–888 (1995).
Dormand, E.-L. & Brand, A. H. Runt determines cell fates in the Drosophila embryonic CNS. Development 125, 1659–1667 ( 1998).
Hacker, U., Lin, X. & Perrimon, N. The Drosophila sugarless gene modulates Wingless signaling and encodes an enzyme involved in polysaccharide biosynthesis. Development 124, 3565–3523 (1997).
Kennerdell, J. R. & Carthew, R. W. Use of dsRNA-mediated genetic interference to demonstrate that frizzled and frizzled 2 act in the Wingless pathway. Cell 95, 1017 –1026 (1998).
Spradling, A. C. in Drosophila, a Practical Approach (ed. Roberts, D. B.) 175–198 (IRL, Oxford, 1986).
Patel, N. H. in Drosophila melanogaster: Practical Uses in Cell and Molecular Biology Vol. 44 (eds Goldstein, L. S. B. & Fyrberg E. A.) 445–487 (Academic, San Diego, 1994).
Acknowledgements
We thank J. Raff and C. Waterman-Storer for discussions; J. Haseloff, J. Raff, H. Chang and G. Rubin for providing DNA constructs, antibodies and/or Drosophila lines; T. Bossing, J. Gray, J. Raff and P. van Roessel for comments on the manuscript; and A. Sossick for technical assistance in video production. J.A.K. is a Wellcome Trust Prize Student. This work was funded by the Wellcome Trust.
Correspondence and requests for materials should be addressed to A.H.B.
Supplementary information is available on Nature Cell Biology’s World-Wide Web site (http://cellbio.nature.com).
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Movie 1 Cell division in epidermoblasts.
In epidermoblasts, the centrosome duplicates on the basal side of the nucleus, and the two centrosomes migrate laterally to opposite sides of the cell. The spindle forms parallel to the surface of the embryo and remains in this orientation throughout mitosis. Microtubules were labelled with tau-GFP, and confocal images were recorded every 12 s. Basal is at the top and apical at the bottom. Lateral view of a stage-10 embryo (also see Fig. 2 of article). (MOV 2702 kb)
Movie 2 Apical centrosome duplication at interphase.
Although the spindle can rotate in either direction, the direction of rotation usually correlates with the position of the centrosome at interphase. When the interphase centrosome duplicates apically, the primed centrosome moves to the more anterior side of the cell before moving apically. The mitotic spindle rotates 90° to a position along the apical-basal axis of the cell. Microtubules were labelled with tau-GFP, and confocal images were recorded every 12 s. Basal is at the top and apical at the bottom. Lateral view of a stage-10 embryo. (MOV 3811 kb)
Movie 3 Basal centrosome duplication at interphase.
Although the spindle can rotate in either direction, the direction of rotation usually correlates with the position of the centrosome at interphase. When the interphase centrosome duplicates basally, the primed centrosome first migrates to the more posterior side of the cell at prophase, and then apically at metaphase. The mitotic spindle rotates 90° to a position along the apical-basal axis of the cell. Microtubules were labelled with tau-GFP and cell cortices with Src-GFP. Images were collected every 20 s. Basal is at the top and apical at the bottom. Lateral view of a stage-10 embryo. (MOV 2490 kb)
Movie 4 The neuroblast spindle rotates 90° at metaphase.
In most cases the spindle is formed before rotation. In this case the bipolar spindle forms as it rotates 90° to its position along the apical-basal axis of the cell. Microtubules were labelled with tau-GFP and cell cortices with Src-GFP. Images were taken every 4 s. Basal is at the top and apical at the bottom. Lateral view of a stage-11 embryo (also see Fig. 2 of article). (MOV 1944 kb)
Movie 5 Loss of Inscuteable impairs spindle rotation.
In embryos injected with inscuteable dsRNA, spindle rotation fails to occur in 10% of neuroblasts. At first the spindle forms normally, perpendicular to the apical-basal axis. The spindle seesaws but does not rotate. In this example, cell division occurs at a 45° angle to the apical-basal axis. Microtubules were labelled with tau-GFP and cell cortices with Src-GFP. Images were taken every 10 s. Basal is at the top and apical at the bottom. Lateral view of a stage-10 embryo (also see Fig. 3 of article). (MOV 2650 kb)
Movie 6
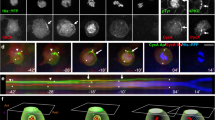
The mitotic spindle becomes asymmetric during neuroblast cell division. The neuroblast mitotic spindle is symmetric and centrally placed at metaphase. At the start of anaphase, the basal cell membrane starts to invaginate and the spindle becomes asymmetric. The distance between the apical and basal centrosome stays the same throughout the division (white arrows mark their starting position and white dots their end point). The midbody, however, gradually moves basally (red arrow), until it is positioned at the site of the cleavage furrow. The neuroblast aster enlarges from anaphase onwards, and the GMC aster is reduced, while the neuroblast half of the spindle elongates and the GMC half becomes shorter. Microtubules were labelled with tau-GFP and cell cortices with Src-GFP. Confocal images were taken every 5 s. Basal is at the top and apical at the bottom. Lateral view of a stage-10 embryo (also see Fig. 4 of article). (MOV 1366 kb)
Movie 7 The mitotic spindle seesaws in epidermoblasts.
The mitotic spindle seesaws in epidermoblasts before undergoing cell division. Microtubules were labelled with tau-GFP, and confocal images were recorded every 20 s. Ventral view of a stage-11 embryo. Anterior is to the left, posterior to the right. (MOV 3857 kb)
Rights and permissions
About this article
Cite this article
Kaltschmidt, J., Davidson, C., Brown, N. et al. Rotation and asymmetry of the mitotic spindle direct asymmetric cell division in the developing central nervous system. Nat Cell Biol 2, 7–12 (2000). https://doi.org/10.1038/71323
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/71323
This article is cited by
-
Long-term in vivo imaging of Drosophila larvae
Nature Protocols (2020)