Abstract
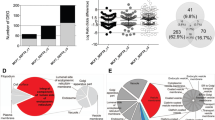
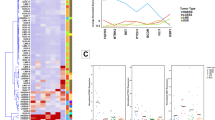
Septins are an evolutionarily conserved family of GTPases with diverse functions including roles in cytokinesis that have been implicated in neoplasia. To address the potential role of SEPT9 in tumorigenesis, we assessed the expression of SEPT9 in 7287 fresh frozen human tissue samples and 292 human cell lines by microarray analysis. In addition, we used a sensitive RT–PCR strategy to define the expression of SEPT9 isoforms in archival formalin-fixed and paraffin-embedded normal human tissues. The mRNA data were further confirmed by immunohistological analyses of SEPT9 protein expression in normal human tissues using antisera that detect SEPT9 isoforms. Using these complementary approaches, we demonstrate that SEPT9 mRNA and protein are expressed ubiquitously, with the isoforms showing tissue-specific expression. The microarray analysis indicates that there is consistent overexpression of SEPT9 in diverse human tumours including breast, CNS, endometrium, kidney, liver, lung, lymphoid, oesophagus, ovary, pancreas, skin, soft tissue and thyroid. Since tumours are commonly associated with enhanced cell proliferation, we examined the possible correlation of Ki67 and SEPT9 expression in normal tissues and tumours. Our data indicate that the overexpression of SEPT9 in neoplasia is not simply a proliferation-associated phenomenon, despite its role in cytokinesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout










Similar content being viewed by others
References
Burrows JF, Chanduloy S, McIlhatton MA, Nagar H, Yeates K, Donaghy P, Price J, Godwin AK, Johnston PG and Russell SE . (2003). J. Pathol., 201, 581–588.
Faty M, Fink M and Barral Y . (2002). Curr. Genet., 41, 123–131.
Field CM and Kellogg D . (1999). Trends Cell Biol., 9, 387–394.
Hall PA and Levison DA . (1990). J. Clin. Pathol., 43, 184–192.
Hall PA and Russell SE . (2004). J. Pathol., 204, 189–505.
Kalikin LM, Sims HL and Petty EM . (2000). Genomics, 63, 165–172.
Kartmann B and Roth D . (2001). J. Cell Sci., 114, 839–844.
Kremer BE and Macara IG . (2003). Mol. Biol. Cell, 14, 447a.
Leipe DD, Wolf YI, Koonin EV and Aravind L . (2002). J Mol. Biol., 317, 41–72.
Martinez C, Sanjuan MA, Dent JA, Karlsson L and Ware J . (2004). Biochem. J., 382, 783–791.
McCormick D, Yu C, Hobbs C and Hall PA . (1993). Histopathology, 22, 543–547.
McIlhatton MA, Burrows JF, Donaghy PG, Chanduloy S, Johnston PG and Russell SE . (2001). Oncogene, 20, 5930–5939.
Montagna C, Lyu MS, Hunter K, Lukes L, Lowther W, Reppert T, Hissong B, Weaver Z and Ried T . (2003). Cancer Res., 63, 2179–2187.
Osaka M, Rowley JD and Zeleznik L . (1999). Proc. Natl. Acad. Sci. USA, 96, 6428–6433.
Peng XR, Jia Z, Zhang Y, Ware J and Trimble WS . (2002). Mol. Cell Biol., 22, 378–387.
Polo S, Pece S and Di Fiore PP . (2004). Curr. Opin. Cell Biol., 16, 156–161.
Robertson C, Church SW, Nagar HA, Price J, Hall PA and Russell SE . (2004). J. Pathol., 203, 519–527.
Russell SE, McIlhatton MA, Burrows JF, Donaghy PG, Chanduloy S, Petty EM, Kalikin LM, Church SW, McIlroy S, Harkin DP, Keilty GW, Cranston AN, Weissenbach J, Hickey I and Johnston PG . (2000). Cancer Res., 60, 4729–4734.
Sager R . (1997). Proc. Natl. Acad. Sci. USA, 94, 952–955.
Sørensen AB, Lund AH, Ethelberg S, Copeland NG, Jenkins NA and Pedersen FS . (2000). J. Virol., 74, 2161–2168.
Sørensen AB, Warming S, Fuchtbauer EM and Pedersen FS . (2002). Gene, 285, 79–89.
Whitfield ML, Sherlock G, Saldanha AJ, Murray JI, Ball CA, Alexander KE, Matese JC, Perou CM, Hurt MM, Brown PO and Botstein D . (2002). Mol. Biol. Cell, 13, 1977–2000.
Acknowledgements
We gratefully acknowledge financial support from the Northern Ireland Research & Development Office, Pathological Society of Great Britain & Ireland and Queen's University Belfast. We thank Adrian Jubb for assistance with data mining and Dr Simon McDade for Figure 1a.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Oncogene website (http://www.nature.com/onc)
Supplementary information
Rights and permissions
About this article
Cite this article
Scott, M., Hyland, P., McGregor, G. et al. Multimodality expression profiling shows SEPT9 to be overexpressed in a wide range of human tumours. Oncogene 24, 4688–4700 (2005). https://doi.org/10.1038/sj.onc.1208574
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1208574
Keywords
This article is cited by
-
Septins, a cytoskeletal protein family, with emerging role in striated muscle
Journal of Muscle Research and Cell Motility (2021)
-
MicroRNA-127-3p regulates myoblast proliferation by targeting Sept7
Biotechnology Letters (2020)
-
Repression of Septin9 and Septin2 suppresses tumor growth of human glioblastoma cells
Cell Death & Disease (2018)
-
Proteomic profile of pre - B2 lymphoblasts from children with acute lymphoblastic leukemia (ALL) in relation with the translocation (12; 21)
Clinical Proteomics (2014)
-
Septin 9 promoter region methylation in free circulating DNA—potential role in noninvasive diagnosis of lung cancer: preliminary report
Medical Oncology (2014)