Abstract
Two-photon laser scanning microscopy of calcium dynamics using fluorescent indicators is a widely used imaging method for large-scale recording of neural activity in vivo. Here, we introduce volumetric two-photon imaging of neurons using stereoscopy (vTwINS), a volumetric calcium imaging method that uses an elongated, V-shaped point spread function to image a 3D brain volume. Single neurons project to spatially displaced 'image pairs' in the resulting 2D image, and the separation distance between projections is proportional to depth in the volume. To demix the fluorescence time series of individual neurons, we introduce a modified orthogonal matching pursuit algorithm that also infers source locations within the 3D volume. We illustrated vTwINS by imaging neural population activity in the mouse primary visual cortex and hippocampus. Our results demonstrated that vTwINS provides an effective method for volumetric two-photon calcium imaging that increases the number of neurons recorded while maintaining a high frame rate.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Denk, W., Strickler, J.H. & Webb, W.W. Two-photon laser scanning fluorescence microscopy. Science 248, 73–76 (1990).
Mank, M. & Griesbeck, O. Genetically encoded calcium indicators. Chem. Rev. 108, 1550–1564 (2008).
Tian, L., Akerboom, J., Schreiter, E.R. & Looger, L.L. Neural activity imaging with genetically encoded calcium indicators. Prog. Brain Res. 196, 79–94 (2012).
Stosiek, C., Garaschuk, O., Holthoff, K. & Konnerth, A. In vivo two-photon calcium imaging of neuronal networks. Proc. Natl. Acad. Sci. USA 100, 7319–7324 (2003).
Dombeck, D.A., Khabbaz, A.N., Collman, F., Adelman, T.L. & Tank, D.W. Imaging large-scale neural activity with cellular resolution in awake, mobile mice. Neuron 56, 43–57 (2007).
Grewe, B.F., Voigt, F.F., van 't Hoff, M. & Helmchen, F. Fast two-layer two-photon imaging of neuronal cell populations using an electrically tunable lens. Biomed. Opt. Express 2, 2035–2046 (2011).
Göbel, W., Kampa, B.M. & Helmchen, F. Imaging cellular network dynamics in three dimensions using fast 3D laser scanning. Nat. Methods 4, 73–79 (2007).
Duemani Reddy, G., Kelleher, K., Fink, R. & Saggau, P. Three-dimensional random access multiphoton microscopy for functional imaging of neuronal activity. Nat. Neurosci. 11, 713–720 (2008).
Kirkby, P.A., Srinivas Nadella, K.M. & Silver, R.A. A compact Acousto-Optic Lens for 2D and 3D femtosecond based 2-photon microscopy. Opt. Express 18, 13721–13745 (2010).
Kong, L. et al. Continuous volumetric imaging via an optical phase-locked ultrasound lens. Nat. Methods 12, 759–762 (2015).
Botcherby, E.J., Juskaitis, R., Booth, M.J. & Wilson, T. An optical technique for remote focusing in microscopy. Opt. Commun. 281, 880–887 (2008).
Chen, T.-W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Botcherby, E.J., Juskaitis, R. & Wilson, T. Scanning two photon fluorescence microscopy with extended depth of field. Opt. Commun. 268, 253–260 (2006).
Thériault, G., Cottet, M., Castonguay, A., McCarthy, N. & De Koninck, Y. Extended two-photon microscopy in live samples with Bessel beams: steadier focus, faster volume scans, and simpler stereoscopic imaging. Front. Cell. Neurosci. 8, 139 (2014).
Lu, R. et al. Video-rate volumetric functional imaging of the brain at synaptic resolution. Nat. Neurosci.http://dx.doi.org/10.1038/nn.4516 (2016).
Mukamel, E.A., Nimmerjahn, A. & Schnitzer, M.J. Automated analysis of cellular signals from large-scale calcium imaging data. Neuron 63, 747–760 (2009).
Maruyama, R. et al. Detecting cells using non-negative matrix factorization on calcium imaging data. Neural Netw. 55, 11–19 (2014).
Pnevmatikakis, E. & Paninski, L. Sparse nonnegative deconvolution for compressive calcium imaging: algorithms and phase transitions. Adv. Neural Inf. Process. Syst. 1250–1258 (2013).
McGloin, D. & Dholakia, K. Bessel beams: diffraction in a new light. Contemp. Phys. 46, 15–28 (2005).
Apthorpe, N.J. et al. Automatic neuron detection in calcium imaging data using convolutional networks. Preprint at http://arxiv.org/abs/1606.07372/ (2016).
Tibshirani, R. Regression shrinkage and selection via the lasso. J. R. Stat. Soc. Series B Stat. Methodol. 58, 267–288 (1996).
Pnevmatikakis, E. et al. A structured matrix factorization framework for large scale calcium imaging data analysis. Preprint at http://arxiv.org/abs/1409.2903/ (2014).
Dombeck, D.A., Harvey, C.D., Tian, L., Looger, L.L. & Tank, D.W. Functional imaging of hippocampal place cells at cellular resolution during virtual navigation. Nat. Neurosci. 13, 1433–1440 (2010).
Yang, W. et al. Simultaneous multi-plane imaging of neural circuits. Neuron 89, 269–284 (2016).
Yaksi, E. & Friedrich, R.W. Recon+struction of firing rate changes across neuronal populations by temporally deconvolved Ca2+ imaging. Nat. Methods 3, 377–383 (2006).
Vogelstein, J.T. et al. Spike inference from calcium imaging using sequential Monte Carlo methods. Biophys. J. 97, 636–655 (2009).
Deneux, T. et al. Accurate spike estimation from noisy calcium signals for ultrafast three-dimensional imaging of large neuronal populations in vivo. Nat. Commun. 7, 12190 (2016).
Oñativia, J., Schultz, S.R. & Dragotti, P.L. A finite rate of innovation algorithm for fast and accurate spike detection from two-photon calcium imaging. J. Neural Eng. 10, 046017 (2013).
Grewe, B.F., Langer, D., Kasper, H., Kampa, B.M. & Helmchen, F. High-speed in vivo calcium imaging reveals neuronal network activity with near-millisecond precision. Nat. Methods 7, 399–405 (2010).
Pachitariu, M. et al. Extracting regions of interest from biological images with convolutional sparse block coding. Adv. Neural Inf. Process. Syst. 24, 1745–1753 (2013).
Pati, Y., Rezaiifar, R. & Krishnaprasad, P. Orthogonal matching pursuit: recursive function approximation with applications to wavelet decomposition. Asilomar Conf. Signals Syst. Comput. 1, 40–44 (1993).
Needell, D. & Tropp, J.A. CoSaMP: iterative signal recovery from incomplete and inaccurate samples. Appl. Comput. Harmon. Anal. 26, 301–321 (2009).
Donoho, D.L., Tsaig, Y., Drori, I. & Starck, J.-L. Sparse solution of underdetermined systems of linear equations by stagewise orthogonal matching pursuit. IEEE Trans. Inf. Theory 58, 1094–1121 (2012).
Prevedel, R. et al. Fast volumetric calcium imaging across multiple cortical layers using sculpted light. Nat. Methods 13, 1021–1028 (2016).
Hopt, A. & Neher, E. Highly nonlinear photodamage in two-photon fluorescence microscopy. Biophys. J. 80, 2029–2036 (2001).
Ji, N., Magee, J.C. & Betzig, E. High-speed, low-photodamage nonlinear imaging using passive pulse splitters. Nat. Methods 5, 197–202 (2008).
Podgorski, K. & Ranganathan, G.N. Brain heating induced by near infrared lasers during multi-photon microscopy. J. Neurophysiol. 116, 1012–1023 (2016).
Kim, C.K. et al. Prolonged, brain-wide expression of nuclear-localized GCaMP3 for functional circuit mapping. Front. Neural Circuits 8, 138 (2014).
Low, R.J., Gu, Y. & Tank, D.W. Cellular resolution optical access to brain regions in fissures: imaging medial prefrontal cortex and grid cells in entorhinal cortex. Proc. Natl. Acad. Sci. USA 111, 18739–18744 (2014).
Cizmár, T. & Dholakia, K. Axial intensity shaping of a Bessel beam. Proc. SPIE 7400, 74001Q (2009).
Watanabe, K. & Microscope Objective Lens, N.C. Japanese Patent no. 2005-189732 (2005).
Madisen, L. et al. Transgenic mice for intersectional targeting of neural sensors and effectors with high specificity and performance. Neuron 85, 942–958 (2015).
Marshel, J.H., Garrett, M.E., Nauhaus, I. & Callaway, E.M. Functional specialization of seven mouse visual cortical areas. Neuron 72, 1040–1054 (2011).
Garrett, M.E., Nauhaus, I., Marshel, J.H. & Callaway, E.M. Topography and areal organization of mouse visual cortex. J. Neurosci. 34, 12587–12600 (2014).
Rickgauer, J.P., Deisseroth, K. & Tank, D.W. Simultaneous cellular-resolution optical perturbation and imaging of place cell firing fields. Nat. Neurosci. 17, 1816–1824 (2014).
Brainard, D.H. The Psychophysics Toolbox. Spat. Vis. 10, 433–436 (1997).
Pelli, D.G. The VideoToolbox software for visual psychophysics: transforming numbers into movies. Spat. Vis. 10, 437–442 (1997).
Kleiner, M., Brainard, D. & Pelli, D.G. What's new in Psychtoolbox-3? Perception 36, 1–16 (2007).
Harvey, C.D., Collman, F., Dombeck, D.A. & Tank, D.W. Intracellular dynamics of hippocampal place cells during virtual navigation. Nature 461, 941–946 (2009).
Domnisoru, C., Kinkhabwala, A.A. & Tank, D.W. Membrane potential dynamics of grid cells. Nature 495, 199–204 (2013).
Aronov, D. & Tank, D.W. Engagement of neural circuits underlying 2D spatial navigation in a rodent virtual reality system. Neuron 84, 442–456 (2014).
Bradski, G. Dr. Dobbs J. Softw. Tools Prof. Program. The OpenCV Library 25, 120,122–125 (2000).
Swirszcz, G., Abe, N. & Lozano, A. Grouped orthogonal matching pursuit for variable selection and prediction. Adv. Neural Inf. Process. Syst. 22, 1150–1158 (2009).
Becker, S., Candes, E. & Grant, M. TFOCS: flexible first-order methods for rank minimization. SIAM Conf. Optim. (2011).
Machado, T.A., Pnevmatikakis, E., Paninski, L., Jessell, T.M. & Miri, A. Primacy of flexor locomotor pattern revealed by ancestral reversion of motor neuron identity. Cell 162, 338–350 (2015).
Harvey, C.D., Coen, P. & Tank, D.W. Choice-specific sequences in parietal cortex during a virtual-navigation decision task. Nature 484, 62–68 (2012).
Acknowledgements
We thank C. Domnisoru, R. Low, and B. Scott for their insightful thoughts and comments. We also thank J. Homann for assistance in using the Psychophysics Toolbox. D.W.T. was supported by NIH grants R01MH083868 and U01NS09054, and the Simons Collaboration on the Global Brain (SCGB 328057). A.C. was supported by an NIH NRSA Training Grant in quantitative neuroscience (T32MH065214). J.W.P. was supported by grants from the McKnight Foundation, the Simons Collaboration on the Global Brain (SCGB 325407), and an NSF CAREER Award (IIS-1150186).
Author information
Authors and Affiliations
Contributions
D.W.T. conceived the project. A.S. and S.Y.T. designed and constructed the vTwINS microscope. S.A.K. and J.L.G. performed the surgery on the mice. A.S. trained the mice and performed the imaging experiments. A.S.C. and J.W.P. designed the SCISM algorithm. A.S.C. implemented SCISM and applied the method to the vTwINS data. A.S. and A.S.C. analyzed the results. A.S., A.S.C., J.W.P., and D.W.T. wrote the manuscript, and all authors provided comments and contributions. J.W.P. and D.W.T. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Simulated elongated point spread functions and alternative vTwINS optical setups.
Simulated 50 μm long PSFs under two sets of Bessel beam parameters (0.34NA left, 0.25NA center) and Gaussian beam parameters (0.175NA right) for the designed microscope setup. The lateral resolution (FWHM) of the Bessel PSFs are 1.12 μm and 1.46 μm, respectively, an improvement over the Gaussian resolution of 2.16 μm. For an equivalent incident power, the relative integrated two-photon excitation of the Bessel beams are 10.3% and 19.3%, respectively, of the Gaussian beam, demonstrating that low-NA Gaussian beams have higher excitation efficiency. (b) Alternate optical setup for simultaneous imaging using vTwINS and conventional TPM. After each frame, a galvanometer switches between a conventional high-NA TPM path and a low-NA Gaussian vTwINS path. A mirror is used to recombine the two paths, with an offset angle of 0.88°. (c) Schematic for alternating single, angled PSFs. A low-NA Gaussian beam is generated with each path before being recombined and imaged onto the scanners. Each scanned frame alternated between each of the two beam paths, corresponding to one half of the vTwINS PSF.
Supplementary Figure 2 vTwINS depth recovery of fluorescent beads.
(a) vTwINS image of a volume containing beads at different locations. Circles depicting locations of all the beads in the vTwINS axial range are color coded by depth (blue is deeper, indicating nearer images, and green is shallower, implying wider images). (b) Spatial profiles recovered automatically from the single vTwINS image are color coded on the same depth scale. Beads at the edge of the field of view have occluded images, and are thus excluded from the analysis as depth cannot be ascertained. (c) Histograms depicting the axial localization error in as well as the total displacement error show that vTwINS can recover bead locations to within approximately 5 μm. (d) The 3D scatter plot compares both the true location of the beads (red dots) and the estimated location (blue circles) to better visualize the vTwINS accuracy.
Supplementary Figure 3 Histogram of maximum spatial overlap estimated from vTwINS profiles for vTwINs PSFs (top) and single axially extended beams (bottom).
(a) Maximum spatial overlap of vTwINS profiles for the V1 dataset. 1.6% of profiles (N=511) have an overlap greater than 50%. (b) Maximum vTwINS overlap with the CA1 dataset. 4.0% of profiles have an overlap greater than 50% (N=882). (c) The left half of vTwINS profiles were taken as equivalent spatial profiles for a single axially extended beam. 18.8% of profiles have a maximum overlap greater than 50% (N=511) for the V1 dataset. (d) 20.5% of single axially extended beam profiles for the CA1 dataset have a spatial overlap greater than 50% (N=882).
Supplementary Figure 4 Matrix factorization on calcium imaging data with high spatial overlap may result in mislabeled profiles.
The alternating beam dataset (Supplementary Fig. 1c) was used to generate a simulated vTwINS dataset. (a) CNMF was run on the right beam of the alternating beam dataset (top trace) and SCISM was run on the simulated vTwINS dataset (lower two traces). The activity from the two cells in the lower traces were merged together when CNMF was run on the same dataset, but with a single axially extended beam. The high spatial overlap between the two profiles resulted in a demixing error. (b) CNMF was run on the simulated vTwINS dataset (top trace) and each of the two single axially extended beams (lower two traces). Profiles from each of the two single axially extended beams that had high spatial overlap with each other were merged together in the simulated vTwINS profile.
Supplementary Figure 5 Sparse convolutional iterative shape matching (SCISM) for demixing vTwINS data.
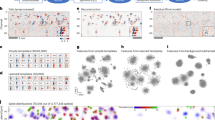
(a) Example stereotyped neuron image pairs. (b) SCISM seeks image pairs at different distances by constructing heat-maps representing the likelihood of a given pair at a given location. Heat maps are calculated by summing the thresholded squared-inner-product between shifts of stereotyped profiles and video frames (shown here with a section of CA1 data). Tλ(·) here denotes the threshold operation. (c) The new spatial profile is chosen at the maximum across all heat maps. (d) The new profile is refined by locally masking and averaging frames closely aligned with the stereotyped spatial profile. (e) The new profile is added to the set of spatial profiles, and the time-traces for all spatial profiles are calculated via non-negative LASSO with sparsity trade-off parameter λ (Methods, Supplementary Note 6). (f) The residual movie is re-computed by subtracting the contribution of the current set of spatial profiles (the sum of outer products of the spatial profiles and their time traces). The algorithm then finds the next spatial profile by iterating from (b) with the new residual.
Supplementary Figure 6 Peak SNR (PSNR) estimates for vTwINS neural profiles for images from V1 (right, n = 511) and CA1 (left, n = 882).
(a) PSNR estimates for full vTwINS profiles (both images used together to estimate time traces). (b) PSNR estimates for the left and right images taken individually. (c) The ratio of PSNR for the full neural profile to the minimum of the PSNR ratios for either the left and right images taken individually. This value is almost universally less than unity, indicating that using the full neural profile (both images simultaneously) is more informative than seeking individual images. The small number of cases where the ratio is above unity indicate singlets where the second image catches a small amount of noise, however these ratios are not so far deviated from unity as to prohibit later processing from finding these singlets.
Supplementary Figure 7 Example vTwINS images from mouse V1 and CA1.
(a) Example frames of full FOV vTwINS data acquired from V1. Top: two examples of vTwINS images from V1. Bottom: Corresponding pre-processed images (5-frame temporal average and two-fold spatial binning) with background-subtraction. Pre-processing makes active pairs of neuronal images more apparent. (b) Example frames of full FOV vTwINS data acquired from CA1. Top: two examples of vTwINS images from V1. Bottom: Corresponding pre-processed images (5-frame temporal average and two-fold spatial binning) with background-subtraction. Pre-processing makes active pairs of neuronal images more apparent. (c) Example frames of half FOV vTwINS data acquired from V1. Top: two examples of vTwINS images from V1. Bottom: Corresponding pre-processed images (5-frame temporal average and two-fold spatial binning) with background-subtraction. Pre-processing makes active pairs of neuronal images more apparent.
Supplementary Figure 8 Example time traces demixed from vTwINS recordings of mouse V1.
(a) 800 second time traces for all 511 demixed V1 spatial profiles. (b) Example time traces for select demixed V1 spatial profiles. SNR and PSNR values are calculated using the 3 σ estimates to obtain the noise standard deviation σ (Supplementary Note 4). The SNR roughly determines the order in which neural profiles are found using SCISM, while the PSNR determines if a neural profile can be found above the noise floor.
Supplementary Figure 9 Anatomical z-stack comparison of found spatial profiles in V1.
(a) Example spatial profiles and time traces from V1. (b) Corresponding location of activity in the anatomical z-stack. White numbers at the top of each image indicates the relative depth in the z-stack.
Supplementary Figure 10 Extended comparison between vTwINS and high-NA spatial profiles and time traces.
(a) Histogram of correlations between vTwINS and high-NA time traces. The majority of correlations cluster tightly around zero, and the tail of outliers indicates the correlations for paired traces. (b) Pearson correlations remain high for the top 98 paired profile traces before falling sharply when no more good pairings remain. The red line indicates the ρ = 0.5 cutoff for determining a pairing (Supplementary Note 4). (c) Four examples of paired traces for both vTwINS imaging (black lines, top of each pair of plots) and high-NA imaging (blue lines, bottom of each pair of plots). Red horizontal lines indicate the 3 σ threshold for significant transients (Supplementary Note 4).
Supplementary Figure 11 Example time traces demixed from vTwINS recordings of mouse CA1.
(a) 1000 second time traces for all 882 demixed CA1 spatial profiles. (b) Example time traces and corresponding SNR and PSNR values for select demixed CA1 spatial profiles. SNR and PSNR values are calculated using the 3 σ estimates to obtain the noise standard deviation σ (Supplementary Note 4). The SNR roughly determines the order in which neural profiles are found using SCISM, while the PSNR determines if a neural profile can be found above the noise floor.
Supplementary Figure 12 Anatomical z-stack comparison of found spatial profiles in CA1.
(a) Example spatial profiles and time traces from CA1. (b) Corresponding location of activity in the anatomical z-stack. White numbers at the top of each image indicates the relative depth in the z-stack.
Supplementary Figure 13 Analysis of alternating-beam variation of vTwINS data taken in mouse CA1.
(a) Found spatial profiles with the left image showing the portion of the profiles from the left beam, and the right image showing the portion of the profiles from the right beam. (b) 100 s of activity for all 1207 found spatial profiles. (c) Example neural profiles and 200 s of activity for the profiles in the subsection of the FOV outlined in white in (a). (d) Histogram of SNR values (N=858 paired) for the neural profiles in the vTwINS equivalent movie, where the left beam and right beam frames are merged together. (e) Histogram of SNR values (Supplementary Note 4) for the spatial profiles in the single-beam frames. (f) Histogram of ratios for the values plotted in (d) and (e) demonstrate that single beam data gains SNR at the cost of having each beam’s data available on only every other frame (half data-rate).
Supplementary Figure 14 Example time traces and corresponding SNR and PSNR values for select demixed spatial profiles from alternating single-beam data from mouse CA1.
SNR and PSNR values are calculated using the 3 σ estimates to obtain the noise standard deviation σ (Supplementary Note 4). The SNR roughly determines the order in which neural profiles are found using SCISM, while the PSNR determines if a neural profile can be found above the noise floor.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–14 and Supplementary Notes 1–6 (PDF 3850 kb)
Supplementary Software
SCISM code and documentation (ZIP 196 kb)
vTwINS V1 example
Video of motion-corrected vTwINS images acquired from V1 (Fig. 4). (AVI 33036 kb)
vTwINS V1 example (background-subtracted)
Video of pre-processed vTwINS images acquired from V1 (Fig. 4). Images were pre-processed images (5-frame temporal average and twofold spatial binning) with background-subtraction (median of movie). (AVI 22572 kb)
vTwINS V1 example
Video of motion-corrected vTwINS images acquired from V1 (Fig. 5). (AVI 24641 kb)
vTwINS V1 example (background-subtracted)
Video of pre-processed vTwINS images acquired from V1 (Fig. 5). Images were pre-processed images (5-frame temporal average and twofold spatial binning) with background-subtraction (median of movie). (AVI 20929 kb)
vTwINS CA1 example
Video of motion-corrected vTwINS images acquired from CA1 (Fig. 6). Images were averaged with 5 frames temporally. (AVI 24719 kb)
vTwINS CA1 example (background-subtracted)
Video of pre-processed vTwINS images acquired from CA1 (Fig. 6). Images were pre-processed images (5-frame temporal average and twofold spatial binning) with background-subtraction (median of movie). (AVI 20126 kb)
SCISM demixing of V1 data
Video corresponding to SCISM demixing for V1 data (Fig. 4). The upper left portion of the videos shows the spatial locations of all profiles found in all iterations thus far. Each pair of circles with a connected line show the pair of images comprising the found profiles. The upper right portion of the video shows the pair locations (circles with connected lines from the upper left portion) super-imposed on the mean-to-variance ration of the movie (a simple measure of activity). This demonstrates that pairs of activity locations f similar strength are picked up. The bottom portion of the video shows the time traces for the set of profiles found at the current iteration. (AVI 3783 kb)
Source data
Rights and permissions
About this article
Cite this article
Song, A., Charles, A., Koay, S. et al. Volumetric two-photon imaging of neurons using stereoscopy (vTwINS). Nat Methods 14, 420–426 (2017). https://doi.org/10.1038/nmeth.4226
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4226
This article is cited by
-
Multifocal fluorescence video-rate imaging of centimetre-wide arbitrarily shaped brain surfaces at micrometric resolution
Nature Biomedical Engineering (2023)
-
Cross-modality supervised image restoration enables nanoscale tracking of synaptic plasticity in living mice
Nature Methods (2023)
-
Fast optical recording of neuronal activity by three-dimensional custom-access serial holography
Nature Methods (2022)
-
Detecting and correcting false transients in calcium imaging
Nature Methods (2022)
-
Non-telecentric two-photon microscopy for 3D random access mesoscale imaging
Nature Communications (2022)