Abstract
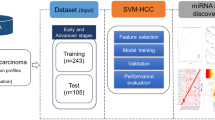
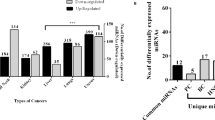
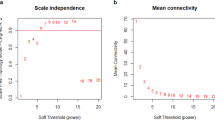
MicroRNAs (miRNAs) are a non-coding family of genes involved in post-transcriptional gene regulation. These transcripts are associated with cell proliferation, cell differentiation, cell death and carcinogenesis. We analysed the miRNA expression profiles in 25 pairs of hepatocellular carcinoma (HCC) and adjacent non-tumorous tissue (NT) and nine additional chronic hepatitis (CH) specimens using a human miRNA microarray. Targets and references samples were co-hybridized to a microarray containing whole human mature and precursor miRNA sequences. Whereas three miRNAs exhibited higher expression in the HCC samples than that in the NT samples, five miRNAs demonstrated lower expression in the HCC samples than in the NT samples (P<0.0001). Classification of samples as HCC or NT by using support vector machine algorithms based on these data provided an overall prediction accuracy of 97.8% (45/46). In addition, the expression levels of four miRNAs were inversely correlated with the degree of HCC differentiation (P<0.01). A comparison of CH and liver cirrhosis samples revealed significantly different pattern of miRNA expression (P<0.01). There were no differences, however, between hepatitis B-positive and hepatitis C-positive samples. This information may help clarify the molecular mechanisms involved in the progression of liver disease, potentially serving as a diagnostic tool of HCC.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bartel DP . (2004). Cell 116: 281–297.
Brechot C, Gozuacik D, Murakami Y, Paterlini-Brechot P . (2000). Semin Cancer Biol 10: 211–231.
Brown MP, Grundy WN, Lin D, Cristianini N, Sugnet CW, Furey TS et al. (2000). Proc Natl Acad Sci USA 97: 262–267.
Calin GA, Dumitru CD, Shimizu M, Bichi R, Zupo S, Noch E et al. (2002). Proc Natl Acad Sci USA 99: 15524–15529.
Esau C, Kang X, Peralta E, Hanson E, Marcusson EG, Ravichandran LV et al. (2004). J Biol Chem 279: 52361–52365.
He L, Hannon GJ . (2004). Nat Rev Genet 5: 522–531.
He L, Thomson JM, Hemann MT, Hernando-Monge E, Mu D, Goodson S et al. (2005). Nature 435: 828–833.
John B, Enright AJ, Aravin A, Tuschl T, Sander C, Marks DS . (2004). PLoS Biol 2: e363.
Johnson SM, Grosshans H, Shingara J, Byrom M, Jarvis R, Cheng A et al. (2005). Cell 120: 635–647.
Ke XS, Liu CM, Liu DP, Liang CC . (2003). Curr Opin Chem Biol 7: 516–523.
Kiriakidou M, Nelson PT, Kouranov A, Fitziev P, Bouyioukos C, Mourelatos Z et al. (2004). Genes Dev 18: 1165–1178.
Liang RQ, Li W, Li Y, Tan CY, Li JX, Jin YX et al. (2005). Nucleic Acids Res 33: e17.
Liu CG, Calin GA, Meloon B, Gamliel N, Sevignani C, Ferracin M et al. (2004). Proc Natl Acad Sci USA 101: 9740–9744.
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D et al. (2005). Nature 435: 834–838.
Michael MZ, O'Connor SM, van Holst Pellekaan NG, Young GP, James RJ . (2003). Mol Cancer Res 1: 882–891.
Moss EG . (2003). Trends Biotechnol 21: 185–187.
Pasquinelli AE, Reinhart BJ, Slack F, Martindale MQ, Kuroda MI, Maller B et al. (2000). Nature 408: 86–89.
Poy MN, Eliasson L, Krutzfeldt J, Kuwajima S, Ma X, Macdonald PE et al. (2004). Nature 432: 226–230.
Takamizawa J, Konishi H, Yanagisawa K, Tomida S, Osada H, Endoh H et al. (2004). Cancer Res 64: 3753–3756.
Th Tsangaris G, Botsonis A, Politis I, Tzortzatou-Stathopoulou F . (2002). Toxicology 178: 135–160.
Thorgeirsson SS, Grisham JW . (2002). Nat Genet 31: 339–346.
van den Berg A, Kroesen BJ, Kooistra K, de Jong D, Briggs J, Blokzijl T et al. (2003). Genes Chromosomes Cancer 37: 20–28.
Xu P, Vernooy SY, Guo M, Hay BA . (2003). Curr Biol 13: 790–795.
Acknowledgements
We thank Itsuro Inoue from the Division of Genetic Diagnosis at the Institute of Medical Science, University of Tokyo, for his helpful advice and comments.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary information accompanies the paper on Oncogene website (http://www.nature.com/onc)
Rights and permissions
About this article
Cite this article
Murakami, Y., Yasuda, T., Saigo, K. et al. Comprehensive analysis of microRNA expression patterns in hepatocellular carcinoma and non-tumorous tissues. Oncogene 25, 2537–2545 (2006). https://doi.org/10.1038/sj.onc.1209283
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1209283
Keywords
This article is cited by
-
MicroRNA Profiling of Root Meristematic Zone in Contrasting Genotypes Reveals Novel Insight into in Rice Response to Water Deficiency
Journal of Plant Growth Regulation (2023)
-
miR-6893-3p is a bonafide negative regulator of splicing activator, RNPS1
3 Biotech (2023)
-
Point-of-care detection assay based on biomarker-imprinted polymer for different cancers: a state-of-the-art review
Polymer Bulletin (2023)
-
The role of miR-153 and related upstream/downstream pathways in cancers: from a potential biomarker to treatment of tumor resistance and a therapeutic target
Medical Oncology (2022)
-
SQSTM1/p62 promotes miR-198 loading into extracellular vesicles and its autophagy-related secretion
Human Cell (2022)